FENYO LAB
Research

Multiomics integration: With the cost of deep sequencing and mass spectrometric analysis decreasing rapidly, it has become feasible to characterize a large number of patient samples by sequencing their genome, transcriptome, proteome, phosphoproteome and metabolome. We have developed proteogenomic integration methods and applied these methods across many cancers to better understand the underlying tumor biology, and to find vulnerabilities that can be exploited in the treatment of patients. (more ...)
Histology and spatial omics: To fully understand biological processes in healthy and diseased tissue, we need to study its spatial structure as it is integral to tissue function. There is a fast improvement in spatial omics technologies that can produce high quality data. We are developing computational methods to better utilize these high dimensional spatial data sets. (more ...)

Cancer biology: With a focus on gynecological and lung cancers, we aim to discover and verify biomarkers and therapeutic targets by integrating multi-modal data—including multi-omics, imaging and electronic health record data—with an ultimate goal of improving patient treatment. (more ...)
Retrotransposons are mobile DNA elements within our genome that can copy and paste themselves into new locations in the genome and can potentially disrupt normal cellular function, and therefore mechanisms suppressing retrotransposition are active in healthy cells, but in many tumors this suppression mechanism is disrupted. We have developed data analysis methods for studying the LINE-1 retrotransposon throughout its lifecycle and we are applying these methods to study the role of retrotransposition in tumors, aging and embryos. (more ...)
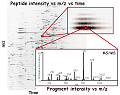
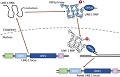

We are developing data analysis methods for genomics for analyzing sequencing data to study replication and quantify retrotransposon expression, proteomics for mass spectrometry data to identify, quantify, characterize and verify proteins and imaging for analyzing super-resolution microscopy and tissue images.
Modeling and simulation of biological systems: We develop models of biological systems and use computer simulations to test the models and compare them to experimental data.